Welcome to the genetic interaction (GI) network data portal where you can access over 88,000 GIs mapped in the HAP1 human haploid cell line. Search your gene of interest to find essentiality profiles, GI scores, GI profiles, and gene co-dependencies.
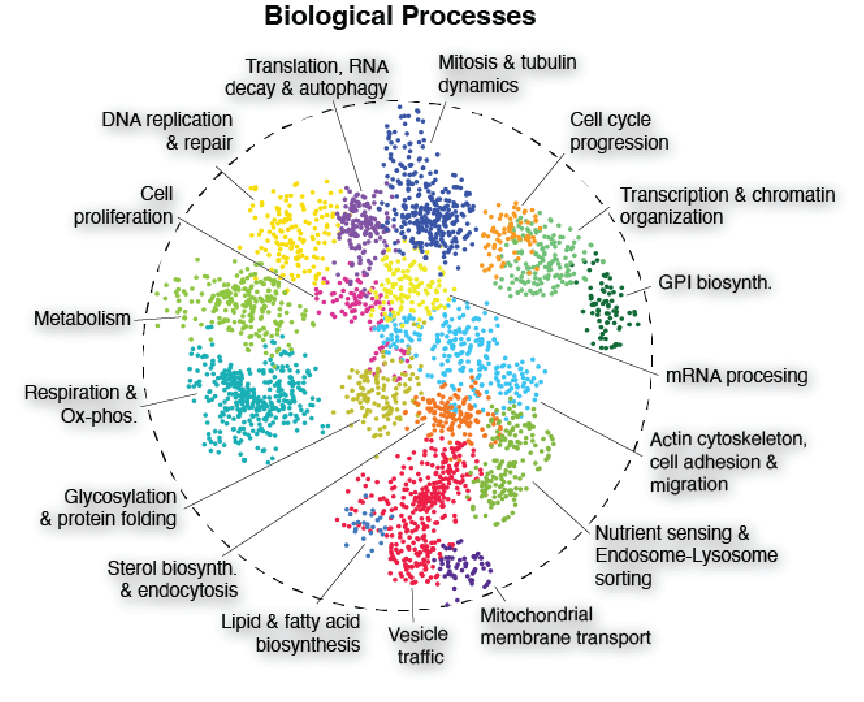
Generated from comprehensive mapping of the human genetic interaction network (GIN) in an isogenic HAP1 cell line background using the state-of-the-art TKOv3 CRISPR library. The current network is derived from >5 million double-mutant fitness measurements including more than 47,000 negative GIs and 41,000 positive GIs.
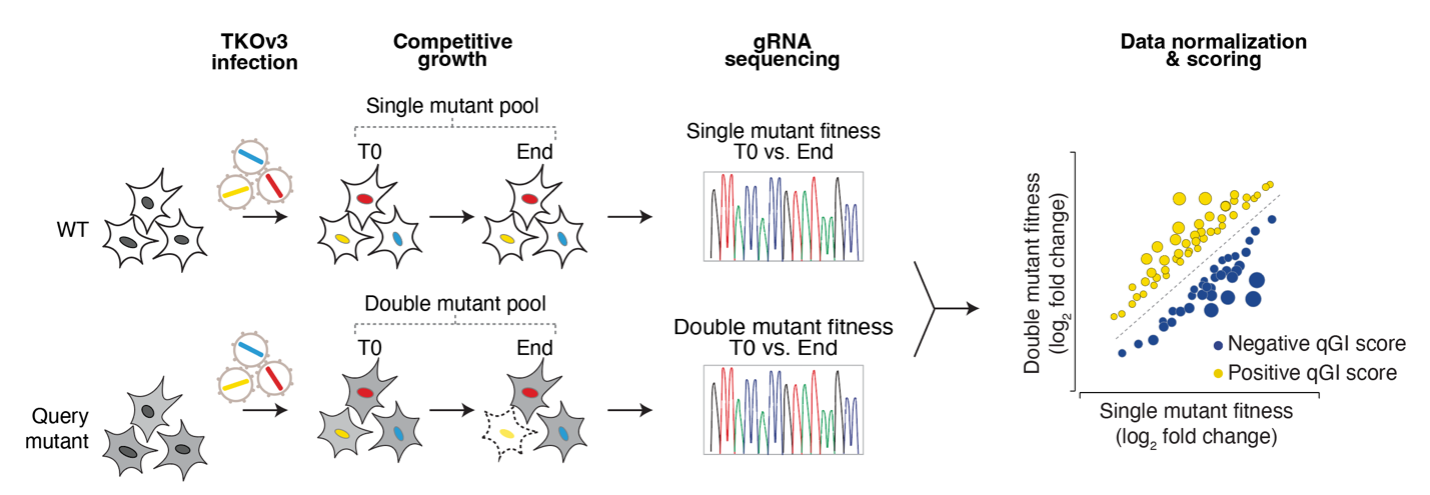
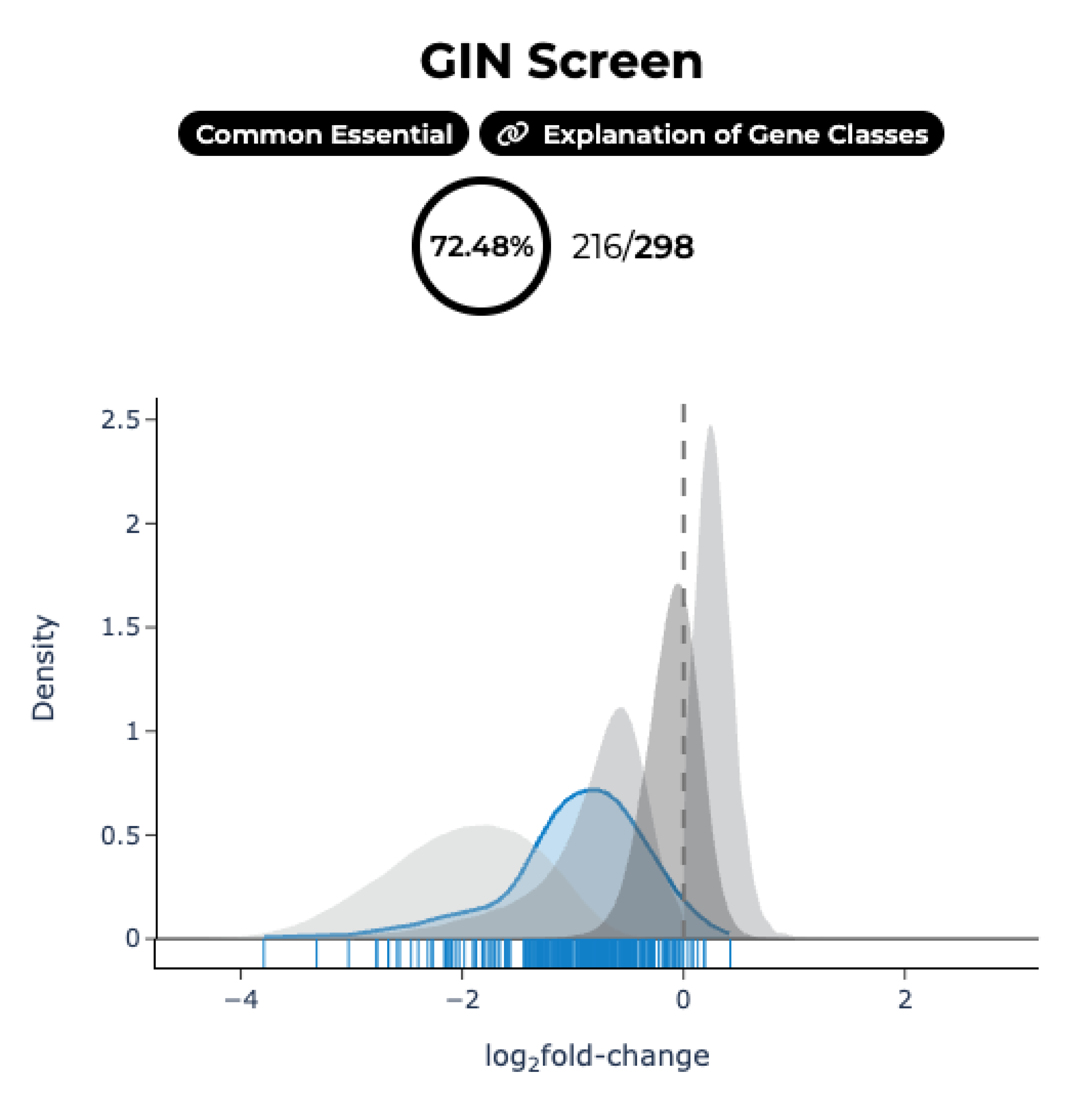
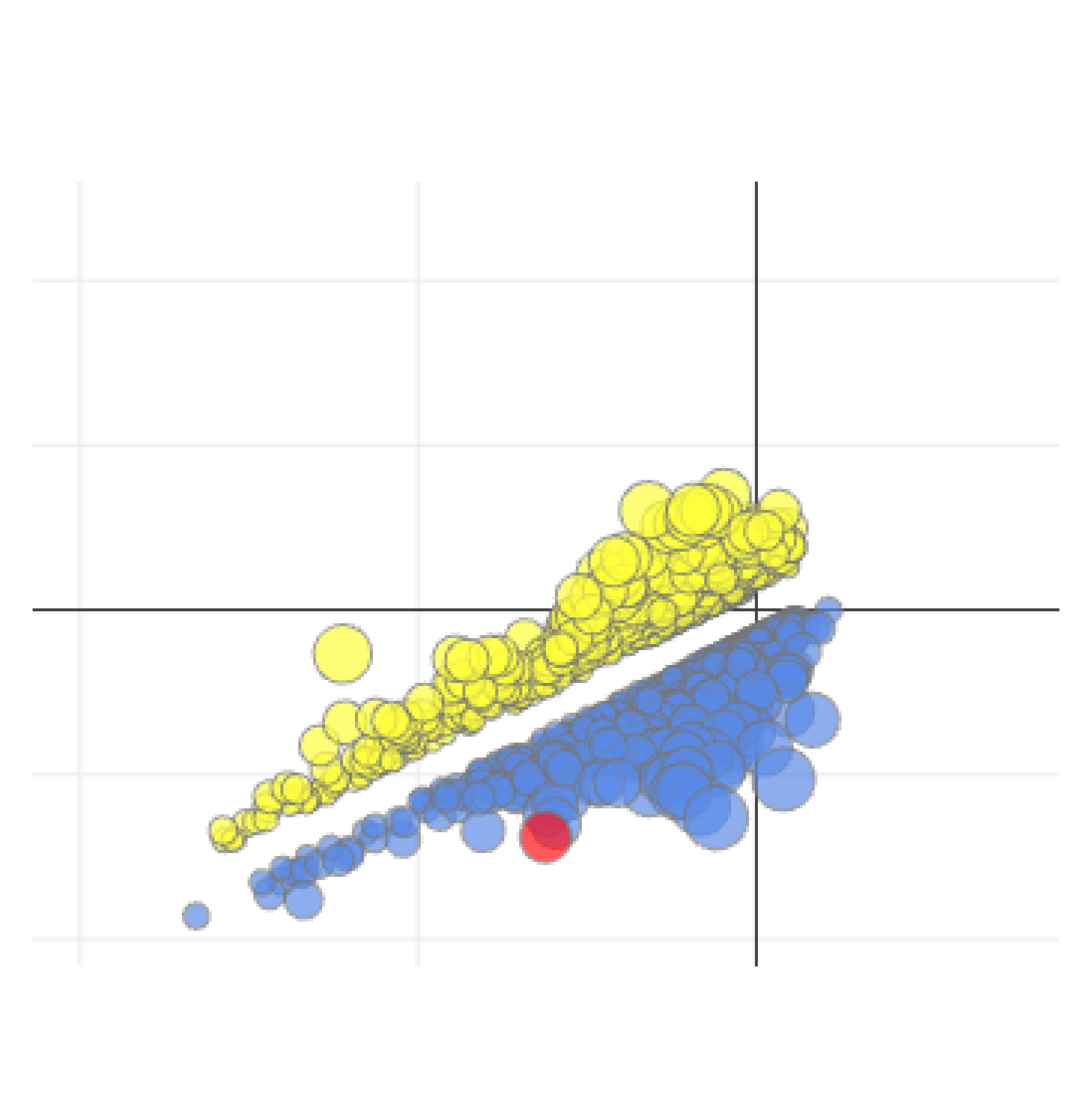
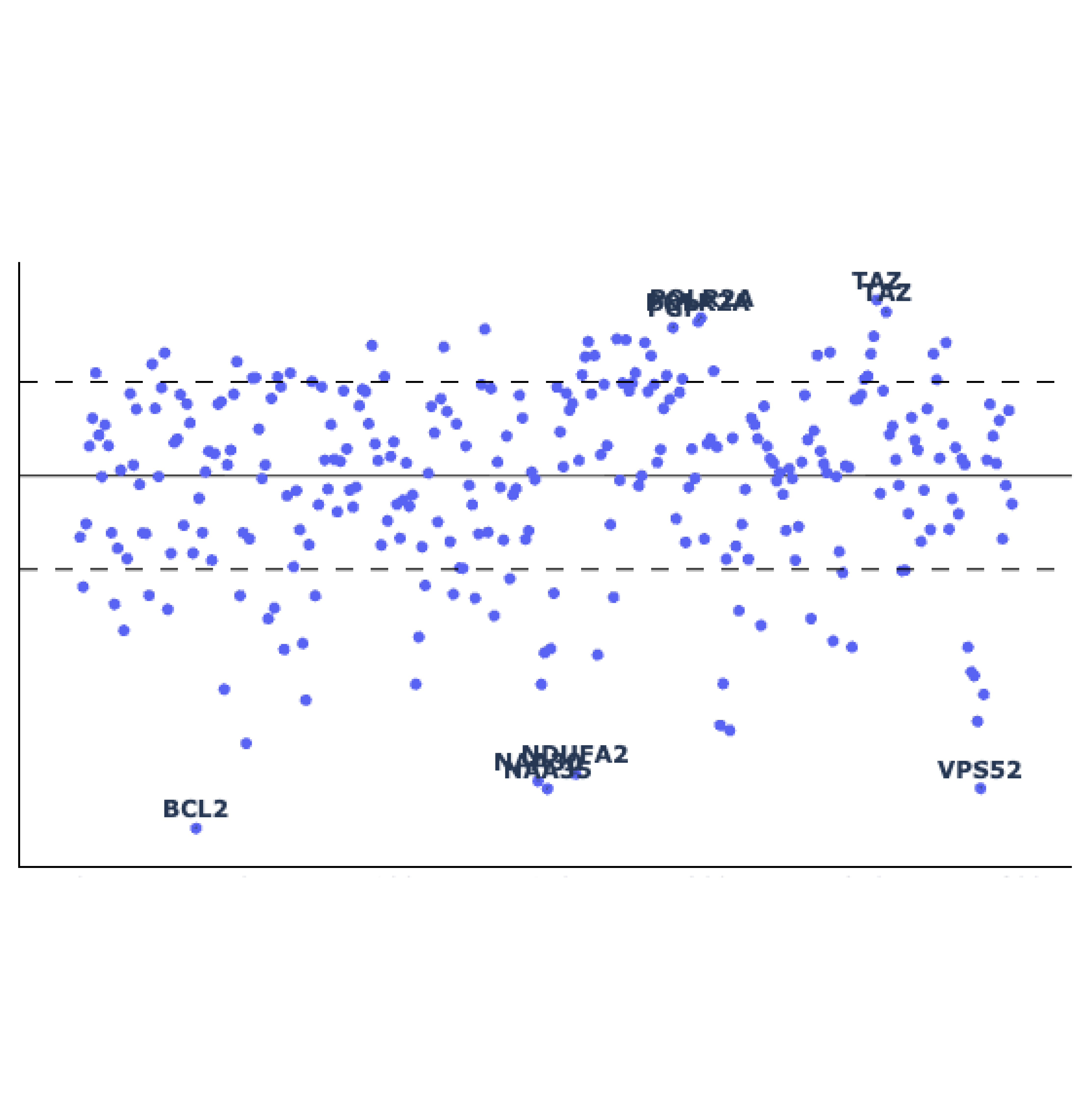
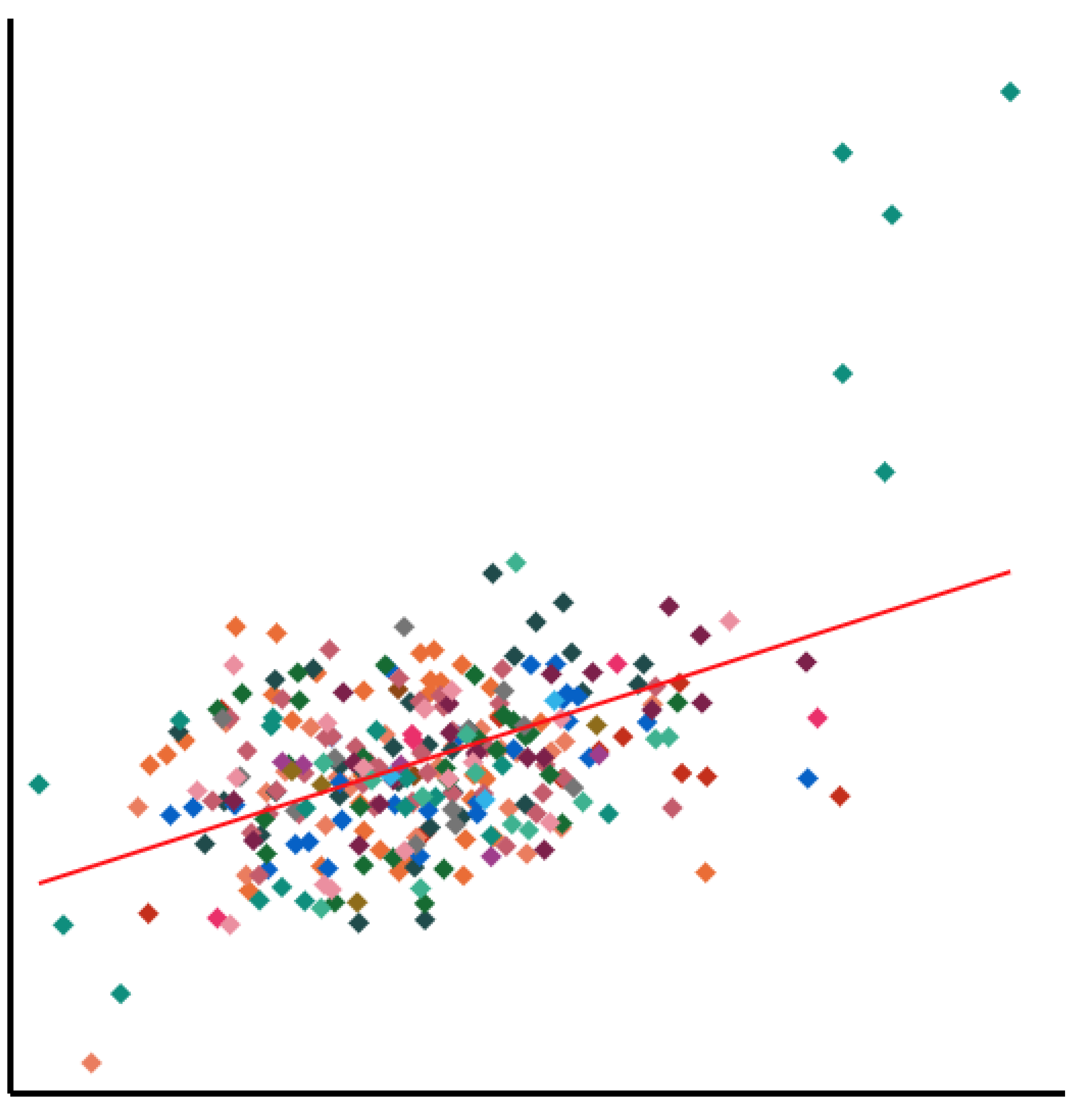
Genome-wide screens are performed in HAP1 cells lacking a given "query" gene. Differential dropout profiles are compared to "hybrid" profiles from 39 wild-type HAP1 screens to identify quantitative genetic interactions (qGI).